
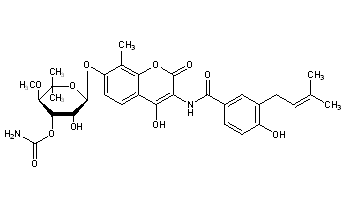
Novobiocin
ノボビオシン;
- Molecular FormulaC31H36N2O11
- Average mass612.624 Da
Monoisotopic: 612.231910004
| INGREDIENT | UNII | CAS | INCHI KEY |
|---|---|---|---|
| Novobiocin sodium | Q9S9NQ5YIY | 1476-53-5 | WWPRGAYLRGSOSU-RNROJPEYSA-M |
Reata Pharmaceuticals Inc
Abgentis is investigating a novobiocin analog, GYR-12 (discovery), as a re-engineered, previously-marketed-but-uncompetitive (undisclosed) antibacterial compound inhibiting ATPase activity of DNA supercoiling GyrB/ParE, for the potential broad-spectrum treatment of bacterial infections, including multi-drug resistant Gram-negative infections. In April 2017, development was underway [1924695].
Novobiocin, also known as albamycin or cathomycin, is an aminocoumarin antibiotic that is produced by the actinomycete Streptomyces niveus, which has recently been identified as a subjective synonym for S. spheroides[1] a member of the order Actinobacteria. Other aminocoumarin antibiotics include clorobiocin and coumermycin A1.[2] Novobiocin was first reported in the mid-1950s (then called streptonivicin).[3][4]
It is active against Staphylococcus epidermidis and may be used to differentiate it from the other coagulase-negative Staphylococcus saprophyticus, which is resistant to novobiocin, in culture.
Novobiocin was licensed for clinical use under the tradename Albamycin (Pharmacia And Upjohn) in the 1960s. Its efficacy has been demonstrated in preclinical and clinical trials.[5][6] The oral form of the drug has since been withdrawn from the market due to lack of efficacy.[7] Novobiocin is an effective antistaphylococcal agent used in the treatment of MRSA.[8]
Mechanism of action
The molecular basis of action of novobiocin, and other related drugs clorobiocin and coumermycin A1 has been examined.[2][9][10][11][12] Aminocoumarins are very potent inhibitors of bacterial DNA gyrase and work by targeting the GyrB subunit of the enzyme involved in energy transduction. Novobiocin as well as the other aminocoumarin antibiotics act as competitive inhibitors of the ATPase reaction catalysed by GyrB. The potency of novobiocin is considerably higher than that of the fluoroquinolones that also target DNA gyrase, but at a different site on the enzyme. The GyrA subunit is involved in the DNA nicking and ligation activity.
Novobiocin has been shown to weakly inhibit the C-terminus of the eukaryotic Hsp90 protein (high micromolar IC50). Modification of the novobiocin scaffold has led to more selective Hsp90 inhibitors.[13] Novobiocin has also been shown to bind and activate the Gram-negative lipopolysaccharide transporter LptBFGC.[14][15]
Structure
Novobiocin is an aminocoumarin. Novobiocin may be divided up into three entities; a benzoic acid derivative, a coumarin residue, and the sugar novobiose.[9] X-ray crystallographic studies have found that the drug-receptor complex of Novobiocin and DNA Gyrase shows that ATP and Novobiocin have overlapping binding sites on the gyrase molecule.[16] The overlap of the coumarin and ATP-binding sites is consistent with aminocoumarins being competitive inhibitors of the ATPase activity.[17]
Structure–activity relationship
In structure activity relationship experiments it was found that removal of the carbamoyl group located on the novobiose sugar lead to a dramatic decrease in inhibitory activity of novobiocin.[17]
Biosynthesis
This aminocoumarin antibiotic consists of three major substituents. The 3-dimethylallyl-4-hydroxybenzoic acid moiety, known as ring A, is derived from prephenate and dimethylallyl pyrophosphate. The aminocoumarin moiety, known as ring B, is derived from L-tyrosine. The final component of novobiocin is the sugar derivative L-noviose, known as ring C, which is derived from glucose-1-phosphate. The biosynthetic gene cluster for novobiocin was identified by Heide and coworkers in 1999 (published 2000) from Streptomyces spheroidesNCIB 11891.[18] They identified 23 putative open reading frames (ORFs) and more than 11 other ORFs that may play a role in novobiocin biosynthesis.
The biosynthesis of ring A (see Fig. 1) begins with prephenate which is a derived from the shikimic acid biosynthetic pathway. The enzyme NovF catalyzes the decarboxylation of prephenate while simultaneously reducing nicotinamide adenine dinucleotide phosphate (NADP+) to produce NADPH. Following this NovQ catalyzes the electrophilic substitution of the phenyl ring with dimethylallyl pyrophosphate (DMAPP) otherwise known as prenylation.[19] DMAPP can come from either the mevalonic acid pathway or the deoxyxylulose biosynthetic pathway. Next the 3-dimethylallyl-4-hydroxybenzoate molecule is subjected to two oxidative decarboxylations by NovR and molecular oxygen.[20] NovR is a non-heme iron oxygenase with a unique bifunctional catalysis. In the first stage both oxygens are incorporated from the molecular oxygen while in the second step only one is incorporated as determined by isotope labeling studies. This completes the formation of ring A.
The biosynthesis of ring B (see Fig. 2) begins with the natural amino acid L-tyrosine. This is then adenylated and thioesterified onto the peptidyl carrier protein (PCP) of NovH by ATPand NovH itself.[21] NovI then further modifies this PCP bound molecule by oxidizing the β-position using NADPH and molecular oxygen. NovJ and NovK form a heterodimer of J2K2 which is the active form of this benzylic oxygenase.[22] This process uses NADP+ as a hydride acceptor in the oxidation of the β-alcohol. This ketone will prefer to exist in its enol tautomer in solution. Next a still unidentified protein catalyzes the selective oxidation of the benzene (as shown in Fig. 2). Upon oxidation this intermediate will spontaneously lactonize to form the aromatic ring B and lose NovH in the process.
The biosynthesis of L-noviose (ring C) is shown in Fig. 3. This process starts from glucose-1-phosphate where NovV takes dTTP and replaces the phosphate group with a dTDP group. NovT then oxidizes the 4-hydroxy group using NAD+. NovT also accomplishes a dehydroxylation of the 6 position of the sugar. NovW then epimerizes the 3 position of the sugar.[23] The methylation of the 5 position is accomplished by NovU and S-adenosyl methionine (SAM). Finally NovS reduces the 4 position again to achieve epimerization of that position from the starting glucose-1-phosphate using NADH.
Rings A, B, and C are coupled together and modified to give the finished novobiocin molecule. Rings A and B are coupled together by the enzyme NovL using ATP to diphosphorylate the carboxylate group of ring A so that the carbonyl can be attacked by the amine group on ring B. The resulting compound is methylated by NovO and SAM prior to glycosylation.[24] NovM adds ring C (L-noviose) to the hydroxyl group derived from tyrosine with the loss of dTDP. Another methylation is accomplished by NovP and SAM at the 4 position of the L-noviose sugar.[25] This methylation allows NovN to carbamylate the 3 position of the sugar as shown in Fig. 4 completing the biosynthesis of novobiocin.
CLIP

CLIP

CLIP

PATENT
US-20190241599
Novel co-crystal forms of novobiocin and its analogs and proline, processes for their preparation and compositions comprising them are claimed. Also claims are methods for inhibiting heat shock protein 90 and treating or preventing neurodegenerative disorders, such as diabetic peripheral neuropathy.
References
- ^ Lanoot B, Vancanneyt M, Cleenwerck I, Wang L, Li W, Liu Z, Swings J (May 2002). “The search for synonyms among streptomycetes by using SDS-PAGE of whole-cell proteins. Emendation of the species Streptomyces aurantiacus, Streptomyces cacaoi subsp. cacaoi, Streptomyces caeruleus and Streptomyces violaceus”. International Journal of Systematic and Evolutionary Microbiology. 52 (Pt 3): 823–9. doi:10.1099/ijs.0.02008-0. PMID 12054245.
- ^ Jump up to:a b Alessandra da Silva Eustáquio (2004) Biosynthesis of aminocoumarin antibiotics in Streptomyces: Generation of structural analogues by genetic engineering and insights into the regulation of antibiotic production. DISSERTATION
- ^ Hoeksema H.; Johnson J. L.; Hinman J. W. (1955). “Structural studies on streptonivicin, a new antibiotic”. J Am Chem Soc. 77 (24): 6710–6711. doi:10.1021/ja01629a129.
- ^ Smith C. G.; Dietz A.; Sokolski W. T.; Savage G. M. (1956). “Streptonivicin, a new antibiotic. I. Discovery and biologic studies”. Antibiotics & Chemotherapy. 6: 135–142.
- ^ Raad I, Darouiche R, Hachem R, Sacilowski M, Bodey GP (November 1995). “Antibiotics and prevention of microbial colonization of catheters”. Antimicrobial Agents and Chemotherapy. 39 (11): 2397–400. doi:10.1128/aac.39.11.2397. PMC 162954. PMID 8585715.
- ^ Raad II, Hachem RY, Abi-Said D, Rolston KV, Whimbey E, Buzaid AC, Legha S (January 1998). “A prospective crossover randomized trial of novobiocin and rifampin prophylaxis for the prevention of intravascular catheter infections in cancer patients treated with interleukin-2”. Cancer. 82 (2): 403–11. doi:10.1002/(SICI)1097-0142(19980115)82:2<412::AID-CNCR22>3.0.CO;2-0. PMID 9445199.
- ^ “Determination That ALBAMYCIN (Novobiocin Sodium) Capsule, 250 Milligrams, Was Withdrawn From Sale for Reasons of Safety or Effectiveness”. The Federal Register. 19 January 2011.
- ^ Walsh TJ, Standiford HC, Reboli AC, John JF, Mulligan ME, Ribner BS, Montgomerie JZ, Goetz MB, Mayhall CG, Rimland D (June 1993). “Randomized double-blinded trial of rifampin with either novobiocin or trimethoprim-sulfamethoxazole against methicillin-resistant Staphylococcus aureus colonization: prevention of antimicrobial resistance and effect of host factors on outcome”. Antimicrobial Agents and Chemotherapy. 37 (6): 1334–42. doi:10.1128/aac.37.6.1334. PMC 187962. PMID 8328783.
- ^ Jump up to:a b Maxwell A (August 1993). “The interaction between coumarin drugs and DNA gyrase”. Molecular Microbiology. 9 (4): 681–6. doi:10.1111/j.1365-2958.1993.tb01728.x. PMID 8231802.
- ^ Maxwell A (February 1999). “DNA gyrase as a drug target”. Biochemical Society Transactions. 27 (2): 48–53. doi:10.1042/bst0270048. PMID 10093705.
- ^ Lewis RJ, Tsai FT, Wigley DB (August 1996). “Molecular mechanisms of drug inhibition of DNA gyrase”. BioEssays. 18 (8): 661–71. doi:10.1002/bies.950180810. PMID 8760340.
- ^ Maxwell A, Lawson DM (2003). “The ATP-binding site of type II topoisomerases as a target for antibacterial drugs”. Current Topics in Medicinal Chemistry. 3 (3): 283–303. doi:10.2174/1568026033452500. PMID 12570764.
- ^ Yu XM, Shen G, Neckers L, Blake H, Holzbeierlein J, Cronk B, Blagg BS (September 2005). “Hsp90 inhibitors identified from a library of novobiocin analogues”. Journal of the American Chemical Society. 127 (37): 12778–9. doi:10.1021/ja0535864. PMID 16159253.
- ^ Mandler MD, Baidin V, Lee J, Pahil KS, Owens TW, Kahne D (June 2018). “Novobiocin Enhances Polymyxin Activity by Stimulating Lipopolysaccharide Transport”. Journal of the American Chemical Society. 140 (22): 6749–6753. doi:10.1021/jacs.8b02283. PMC 5990483. PMID 29746111.
- ^ May JM, Owens TW, Mandler MD, Simpson BW, Lazarus MB, Sherman DJ, Davis RM, Okuda S, Massefski W, Ruiz N, Kahne D (December 2017). “The Antibiotic Novobiocin Binds and Activates the ATPase That Powers Lipopolysaccharide Transport”. Journal of the American Chemical Society. 139 (48): 17221–17224. doi:10.1021/jacs.7b07736. PMC 5735422. PMID 29135241.
- ^ Tsai FT, Singh OM, Skarzynski T, Wonacott AJ, Weston S, Tucker A, Pauptit RA, Breeze AL, Poyser JP, O’Brien R, Ladbury JE, Wigley DB (May 1997). “The high-resolution crystal structure of a 24-kDa gyrase B fragment from E. coli complexed with one of the most potent coumarin inhibitors, clorobiocin”. Proteins. 28 (1): 41–52. doi:10.1002/(sici)1097-0134(199705)28:1<41::aid-prot4>3.3.co;2-b. PMID 9144789.
- ^ Jump up to:a b Flatman RH, Eustaquio A, Li SM, Heide L, Maxwell A (April 2006). “Structure-activity relationships of aminocoumarin-type gyrase and topoisomerase IV inhibitors obtained by combinatorial biosynthesis”. Antimicrobial Agents and Chemotherapy. 50 (4): 1136–42. doi:10.1128/AAC.50.4.1136-1142.2006. PMC 1426943. PMID 16569821.
- ^ Steffensky M, Mühlenweg A, Wang ZX, Li SM, Heide L (May 2000). “Identification of the novobiocin biosynthetic gene cluster of Streptomyces spheroides NCIB 11891”. Antimicrobial Agents and Chemotherapy. 44 (5): 1214–22. doi:10.1128/AAC.44.5.1214-1222.2000. PMC 89847. PMID 10770754.
- ^ Pojer F, Wemakor E, Kammerer B, Chen H, Walsh CT, Li SM, Heide L (March 2003). “CloQ, a prenyltransferase involved in clorobiocin biosynthesis”. Proceedings of the National Academy of Sciences of the United States of America. 100 (5): 2316–21. Bibcode:2003PNAS..100.2316P. doi:10.1073/pnas.0337708100. PMC 151338. PMID 12618544.
- ^ Pojer F, Kahlich R, Kammerer B, Li SM, Heide L (August 2003). “CloR, a bifunctional non-heme iron oxygenase involved in clorobiocin biosynthesis”. The Journal of Biological Chemistry. 278 (33): 30661–8. doi:10.1074/jbc.M303190200. PMID 12777382.
- ^ Chen H, Walsh CT (April 2001). “Coumarin formation in novobiocin biosynthesis: beta-hydroxylation of the aminoacyl enzyme tyrosyl-S-NovH by a cytochrome P450 NovI”. Chemistry & Biology. 8 (4): 301–12. doi:10.1016/S1074-5521(01)00009-6. PMID 11325587.
- ^ Pacholec M, Hillson NJ, Walsh CT (September 2005). “NovJ/NovK catalyze benzylic oxidation of a beta-hydroxyl tyrosyl-S-pantetheinyl enzyme during aminocoumarin ring formation in novobiocin biosynthesis”. Biochemistry. 44 (38): 12819–26. CiteSeerX 10.1.1.569.1481. doi:10.1021/bi051297m. PMID 16171397.
- ^ Thuy TT, Lee HC, Kim CG, Heide L, Sohng JK (April 2005). “Functional characterizations of novWUS involved in novobiocin biosynthesis from Streptomyces spheroides”. Archives of Biochemistry and Biophysics. 436 (1): 161–7. doi:10.1016/j.abb.2005.01.012. PMID 15752721.
- ^ Pacholec M, Tao J, Walsh CT (November 2005). “CouO and NovO: C-methyltransferases for tailoring the aminocoumarin scaffold in coumermycin and novobiocin antibiotic biosynthesis”. Biochemistry. 44 (45): 14969–76. doi:10.1021/bi051599o. PMID 16274243.
- ^ Freel Meyers CL, Oberthür M, Xu H, Heide L, Kahne D, Walsh CT (January 2004). “Characterization of NovP and NovN: completion of novobiocin biosynthesis by sequential tailoring of the noviosyl ring”. Angewandte Chemie. 43 (1): 67–70. doi:10.1002/anie.200352626. PMID 14694473.
External links
- Novobiocin bound to proteins in the PDB
 |
|
 |
|
| Clinical data | |
|---|---|
| AHFS/Drugs.com | International Drug Names |
| Routes of administration |
intravenous |
| ATCvet code | |
| Pharmacokinetic data | |
| Bioavailability | negligible oral bioavailability |
| Metabolism | excreted unchanged |
| Elimination half-life | 6 hours |
| Excretion | renal |
| Identifiers | |
| CAS Number | |
| PubChem CID | |
| DrugBank | |
| ChemSpider | |
| UNII | |
| KEGG | |
| ChEMBL | |
| CompTox Dashboard(EPA) | |
| ECHA InfoCard | 100.005.589 |
| Chemical and physical data | |
| Formula | C31H36N2O11 |
| Molar mass | 612.624 g·mol−1 |
| 3D model (JSmol) | |
4309-70-0 CAS
calcium;7-[(2R,3R,4S,5R)-4-carbamoyloxy-3-hydroxy-5-methoxy-6,6-dimethyloxan-2-yl]oxy-3-[[4-hydroxy-3-(3-methylbut-2-enyl)benzoyl]amino]-8-methyl-2-oxochromen-4-olate
///////// Novobiocin, ノボビオシン , Antibacterial, Antimicrobial, crystallinic acid, streptonivicin,
History
Novobiocin is a coumarin antibiotic obtained from Streptomyces niveus and other Streptomyces species. Novobiocin is useful primarily in infections involving staphylococci, and other gram-positive organisms. It acts by inhibiting the initiation of DNA replication in bacterial and mammanlian cells. Evidences indicated that Novobiocin blocks prokaryotic DNA gyrase and eukaryotic II topoisomerase, enzymes that relax super-coiled DNA and are crucial for DNA replication.1
Novobiocin
| UIPAC Name | 4-Hydroxy-3-4-hydroxy-3-(3-methylbut-2-enyl)benzamido-8-methylcoumarin-7-yl 3-O-carbamoyl-5,5-di-C-methyl-α-l-lyxofuranoside |
| CAS Number | 303-81-1 |
| Molecular Mass | 612.624 g / mol |
| Chemical Formular | C31H36N2O11 |
Biosynthesis
The substituted coumarin (ring B, red) and the 4-OH benzoyl moiety (ring A, aqua) in novobiocin were derived from -Tyr based on earlier labeling studies. β-OH-Tyr is proposed to be a common intermediate in these two biosynthetic pathways.2

NovH is a -Tyr specific didomain NRPS that generates the
-tyrosyl-S-NovH intermediate. NovH, isolated from E. coli is primed by a PPTase with CoA. The A domain activates
-Tyr as
-tyrosyl-AMP and then transfers the
-tyrosyl group to the HS-pant-PCP domain of NovH through thioester formation.3

-tyrosyl-S-NovH is then function as a cytochrome P450 monooxygenase that hydroxylates the β-carbon of the tethered
-tyrosyl group on NovH. While the substrate
-tyrosyl-S-NovH provides two electrons for a single round of the hydroxylation reaction, the other two electrons needed to reduce the oxygen atom are provided by NADPH via two-electron transfer effected by electron transfer proteins ferrodoxin (Fd) and ferrodoxin reductase (Fd Red).3 The electron transfer route is from NADPH→FAD in Fd Red→Fe–S center in Fd→Heme in NovI→oxygen.

Both NovJ and NovK are similar to 3-keto-ACP reductase and they may form a heterodimer and operate in the reverse direction to oxidize 3-OH to 3-keto. NovO is similar to some quinone C-methyltransferases 3 but the timing of methylation is not clear. NovC resembles flavin-dependent monooxygenases (35 and 32% similarity to dimethylaniline and cyclohexanone monooxygenases, respectively) 3 and is proposed to hydroxylate the ortho position of the phenyl ring. The nucleophilic attack of the ortho hydroxyl group on the thioester carbonyl center would release the coumarin ring and regenerate NovH. Ring B is then synthesized.

Synthesis




Mechanism of action
E.Coli DNA gyrase utilizes ATP to catalyze the negative supercoiling, or under-twisting, of duplex DNA. The energy coupling components of the supercoiling reaction includes 1) the DNA-dependent hydrolysis that converts ATP to ADP and Pi, and 2) the gyrase cleavage reaction that targets the specified DNA site. The two activities are induced by treating the stable gyrase-DNA complex trapped by the inihibitor oxolinic acid with sodium dodecyl sulfate (SDS or Sulphate). 4 Novobiocin competes with ATP in the ATPase and supercoiling assays, hence Novobiocin prevents the ATP from shifting the primary cleavage site on ColE1 DNA by places the site of action of the antibiotics at a reaction step prior to ATP hydrolysis and blocks the binding of ATP. 4 Such a simple mechanism of action represents for all effects of the drugs on DNA gyrase.
Clinical Use
Due to factors as low solubility, poor pharmacokinetics, and limited activity agasinst Gram-negative bacteria, the clinical usage of Novobiocin is not achieved. 5 Therefore, it is of interest to study the novobiocin biosynthetic pathway in order to generate analogs with enhanced solubility and pharmacokinetic properties while maintaining the gyrase inhibitory properties.
References
1 J.C. D’Halluin, M. Milleville, and P. Boulanger. “Effect of Novobiocin on adenovirus DNA synthesis and encapsidation”. Nucleic Acids Research 1980; 8: 1625-1641
2 M. Steffensky, S.M. Li and L. Heide, “Cloning, overexpression, and purification of novobiocic acid synthetase from Streptomyces spheroides ” NCIB 11891. J. Biol. Chem. 275 (2000), pp. 21754–21760.
3 Huawei Chen and Christopher T. Walsh, “Coumarin formation in novobiocin biosynthesis: β-hydroxylation of the aminoacyl enzyme tyrosyl-S-NovH by a cytochrome P450 NovI” Chemistry and Biology; 2001; 8: 301-312
4 K. Scheirer and N. P. Higgins. “The DAN Cleavage Reaction of DNA Gyrase ” The Journal of Biological Chemistry; 1997; 272 (43): 27202-27209
5 N Pi, C. L. F. Meyers, M. Pacholec, C. T. Walsh, and J. A. Leary. “Mass spectrometric characterization of a three-enzyme tandem reacton for assembly and modification of the novobiocin skeleton” PNAS 2004;101;10036-10041










